The Xavante Indians and Genetic Divergence Between American Indians and Africans
PLoS ONE 7 (8): e42702. doi:10.1371/journal.pone.0042702
Genome-Wide Analysis in Brazilian Xavante Indians Reveals Low Degree of Admixture
Patricia C. Kuhn, Andréa R. V. Russo. Horimoto, José Maurício Sanches, João Paulo B. Vieira Filho, Luciana Franco, Amaury Dal Fabbro, Laercio Joel Franco, Alexandre C. Pereira, Regina S. Moises
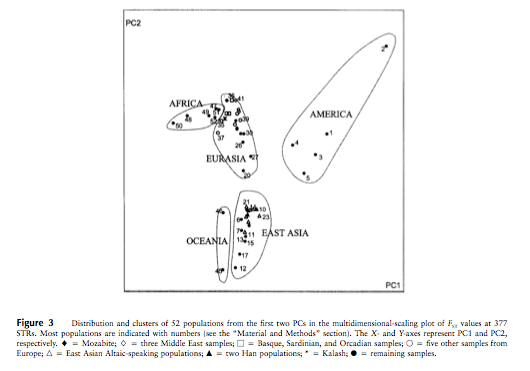
Characterization of population genetic variation and structure can be used as tools for research in human genetics and population isolates are of great interest. The aim of the present study was to characterize the genetic structure of Xavante Indians and compare it with other populations. The Xavante, an indigenous population living in Brazilian Central Plateau, is one of the largest native groups in Brazil. A subset of 53 unrelated subjects was selected from the initial sample of 300 Xavante Indians. Using 86,197 markers, Xavante were compared with all populations of HapMap Phase III and HGDP-CEPH projects and with a Southeast Brazilian population sample to establish its population structure. Principal Components Analysis showed that the Xavante Indians are concentrated in the Amerindian axis near other populations of known Amerindian ancestry such as Karitiana, Pima, Surui and Maya and a low degree of genetic admixture was observed. This is consistent with the historical records of bottlenecks experience and cultural isolation. By calculating pair-wise Fst statistics we characterized the genetic differentiation between Xavante Indians and representative populations of the HapMap and from HGDP-CEPH project. We found that the genetic differentiation between Xavante Indians and populations of Amerindian, Asian, European, and African ancestry increased progressively. Our results indicate that the Xavante is a population that remained genetically isolated over the past decades and can offer advantages for genome-wide mapping studies of inherited disorders.
Link (Free Access PDF)
This genome-wide study of an unadmixed Amazonian population speaking a Ge language continues a long-standing tradition of genetic research among isolated South American tribes. The importance of small-deme studies in South America (in addition to classic Karitiana and Surui we now have Xavante) is that they “provide a model of [modern human] ancestral population, because they perhaps maintain similar lifestyle and population size.” (Zhivotovsky, Lev A., Noah A. Rosenberg, and Marcus W. Feldman. 2003. “Features of Evolution and Expansion of Modern Humans, Inferred from Genomewide Microsatellite Markers,” Am J Hum Genet 72 (5), 1178). Small population size keeps variance growth in check, so that
“a size of 700 for the ancient ancestral population would produce an expected value of variance in the number of repeats of approximately 2×700×0.00064=0.90, which is smaller than the average variance at the 271 tetranucleotide loci in Karitiana and Surui, 1.8. This suggests that the variance in repeat scores in an ancestral population, V0, was smaller than that in these American populations and provides an additional argument for using variances in these populations as an upper bound for V0 in dating the first population split in the deep history of modern humans.” (Zhivotovsky et al. 2003, 1181).
Xavante are not that small – they number 10,000 people, which is still smaller than geneticists’ darlings, African foragers – !Kung and Biaka, – but not nearly close to the population size of Karitiana and Surui. Most importantly, Xavante seem to have been genuinely isolated, unlike !Kung or Biaka Pygmies. Producing a genetic study of an isolated tribe is like extracting ancient DNA from a Pleistocene sample. The challenges are different but still formidable. Kuhn et al. (2012) had to take an extra precaution and purge their sample of 300 Xavante Indians of all cryptic close consanguinity. They used ML-Relate software to screen out pairs of parent-offspring, full-siblings and half-siblings. After that the individual with the greatest number of relationships was identified and removed from the Xavante sample. Next, the relatedness was recalculated, generating other relationship structures. Again, the individual with the higher number of relatedness was removed and the relatedness was recalculated. This process was repeatedly made until we had only an unrelated sample. They ended up with 1/6 of their original Xavante sample, or with 53 individuals.
This cleanup of consanguinity left intact any identity-by-descent (IBD) between first-cousins, second-cousins and uncle-nieces. We know from ethnographic sources that Xavante prefer marrying a cross-cousin but, unlike Surui, not a bilateral cross-cousin. Marriages with sister’s daughter are again, unlike Suirui, not practiced. The level of innbreeding, therefore, is expected to be quite high in any Xavante sample, but not as high as in a Surui sample.
Small, isolated demes typically show reduced levels of allelic and locus diversity, higher homozygosity and higher linkage disequilibrium. This was all observed in Xavante, which makes them an ideal candidate for identification of genetic susceptibility for monogenic and polygenic diseases. In Africa, the closest approximation to the South American situation are not !Kung or Biaka Pygmies but Hadza, which are also suspected to have high degree of consanguinity (Lachance et al. 2012. “Evolutionary History and Adaptation from High-Coverage Whole-Genome Sequences of Diverse African Hunter-Gatherers,” Cell 150 (1-3), 5). I blogged about it here.
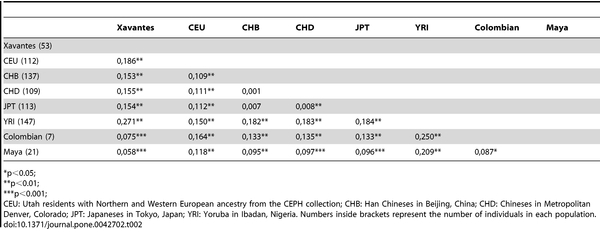
Kuhn et al. (2012) confirm earlier studies that showed that South American Indians and Sub-Saharan Africans are the most divergent populations in the world. Below is the table pooling Fst statistics from their worldwide sample showing the highest value (0.271) separating Xavante and Yoruba.