Neandertal Admixture in Africans: A Back-Migration to Sub-Saharan Africa Confirmed
Genome Biology and Evolution 2013 Oct 25.
Apparent Variation in Neanderthal Admixture among African Populations is Consistent with Gene Flow from non-African Populations.
Shuoguo Wang, Joseph Lachance, Sarah Tishkoff, Jody Hey, and Jinchuan Xing
Abstract
Recent studies have found evidence of introgression from Neanderthals into modern humans outside of sub-Saharan Africa. Given the geographic range of Neanderthals, the findings have been interpreted as evidence of gene exchange between Neanderthals and the modern humans descended from the Out-of-Africa (OOA) migration. Here we examine an alternative interpretation in which the introgression occurred earlier within Africa, between ancestors or relatives of Neanderthals and a subset of African modern humans who were the ancestors of those involved in the OOA migration. Under the alternative model, if the population structure among present-day Africans predates the OOA migration, we might find some African populations show a signal of Neanderthal introgression while others do not. To test this alternative model we compiled a whole-genome data set including 38 sub-Saharan Africans from eight populations and 25 non-African individuals from five populations. We assessed differences in the amount of Neanderthal-like SNP alleles among these populations and observed up to 1.5% difference in the number of Neanderthal-like alleles among African populations. Further analyses suggest that these differences are likely due to recent non-African admixture in these populations. After accounting for recent non-African admixture, our results do not support the alternative model of older (e.g., >100 kya) admixture between modern human and Neanderthal-like hominid within Africa.
The paper confirms the presence of Neandertal admixture in Sub-Saharan African populations and provides evidence that this admixture affected mostly East African populations (Hadza, Sandawe, Maasai). The authors tested whether this admixture occurred between an African ancestor of Neandertals and Africans prior to the putative out-of-Africa event or between Neandertal-admixed (Green et al. 2010) modern non-Africans and modern Africans. They concluded that Neandertal alleles detected in East African populations derive from a back-migration of modern non-Africans to Africa.
Below on the left we can see the four-way comparison values from Wang et al. 2013 (San, H2-African, Neandertal, Chimpanzee). On the right we see the four-way values from the seminal paper by Green et al. 2010 ascertaining “Neandertal admixture” in non-Africans (San, H2-non-African, Neandertal, Chimpanzee). If, for simplicity, we limit our inspection to only the lines that have San as H1, we can see that the highest African values (African Americans, ASW, Hadza, Sandawe and Maasai) and the lowest non-African values (Papuan and French) come somewhat close to each other, while Pygmies and Han are the furthest apart (D = -0.69 vs. -4.8, respectively).
Wang et al. 2013 conclude that Pygmies and San Bushmen were not affected by this wave of returning ex-Africans. Technically they were, but statistically the values resulting from comparison of Pygmy and San genomes with the Neandertal genome are insignificant.
Wang et al. (2013) show strange reluctance to accept, without caveats, an earlier finding that Asians are more Neandertal-admixed than Europeans. They write,
“Another prediction of model ii is that the initial Eurasian populations derived from the OOA migration carried similar amounts of Neanderthal-like alleles. This prediction seemingly contradicts the observation that modern East Asians share more alleles with Neanderthal than modern Europeans. Nevertheless, several factors, including ascertainment bias, genetic drift, and admixture with other archaic human forms, could contribute to the observed difference (Keinan, et al. 2007; Skoglund and Jakobsson 2011). In some situations, a combination of ascertainment bias and genetic drift could generate artificial differences between populations that had the same admixture history (Skoglund and Jakobsson 2011).”
It seems that the same factors (genetic drift + ascertainment bias) could generate the result they obtained for Africa but for some reason they choose apply a double standard and to cast doubt on the varying degree of Neandertal admixture outside of Africa but not in Africa. If they accepted the former finding, then they would have seen a noticeable cline between strongly Neandertal-admixed Asians through moderately Neandertal-admixed Europeans and North Africans to lowly Neandertal-admixed (East) Africans. (Lower levels of Neandertal admixture among Papuans likely reflect an overlay of Denisovan genes.) The usual question of why Asians are more Neandertaloid than Europeans should be reformulated as the question of why Asians are more Neandertaloid than Europeans and East Africans.
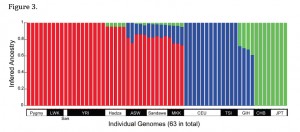
Another intriguing fact to be unearthed from Wang et al. 2013 is that Hadza has the least amount of Neandertal ancestry in East Africa (Sandawe and Maasai are upwards of 16% non-African, Wang et al. 2013, p. 7), while it is more Asian-admixed (green bars on the image on the left) than the other two East African populations (Sandawe and Maasai), according to ADMIXTURE. ADMIXTURE results closely parallel D-statistics (Wang et al. 2013, p. 7), hence, assuming that Asians are the source of Neandertal admixture in Europeans and Africans, it looks like Asians had brought Neandertal genes to Hadza before Europeans brought Neandertal genes (and some Asian genes) to Maasai and Sandawe.
We need to remind ourselves that Hadza are genetically unique in Sub-Saharan Africa by being genetically homogeneous and having high levels of linkage disequilibrium (LD) and long runs of homozygosity – all those molecular traits typically found in such geographic isolates as Amerindians, Papuans or Denisovans (see here). From this perspective, the very presence of Neandertal admixture in Hadza is puzzling because this should have made them equally – if not more – diverse than the rest of Africans. What Wang et al. 2013 do not consider is the possibility that San and Pygmies are not Neandertal-admixed because they have genes introgressed from another, African-specific, hominin (for Khoisans, see here). If this possibility is considered, then the presence of Neandertal-admixture in East Africa marks the colonization of Africa by an Asian population, while the divergence of San and Pygmies and their greater number of chimpanzee genomic matches compared to the rest of modern humans can be explained as the influence of an archaic African substrate and not as a sign of their basal position on a modern human tree.
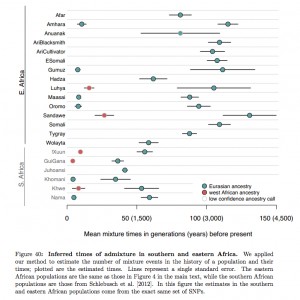 But the very fact that Wang et al. (2013) failed to detect a statistically significant fraction of Neandertal ancestry in San Bushmen is puzzling in and of itself considering that another recent study (not cited in Wang et al. 2013) reported West Eurasian admixture can be found all over South African foragers and pastoralists due to a pastoral migration from East Africa to South Africa in the past 3,000 years (see on the left, from Pickrell, Joseph K., Nick Patterson, Po-Ru Loh, Mark Lipson, Bonnie Berger, Mark Stoneking, Brigitte Pakendorf, and David Reich. 2013. “Ancient West Eurasian Ancestry in Southern and Eastern Africa”). Considering that ancient admixture follows the matreshka principle (“It should be noted that although ADMIXTURE predicts the contribution of three putative ancestral populations in each individual genome, the contribution could be indirect…MKK and Sandawe might not have large amounts of Eurasian ancestry. They have large amounts of Cushitic ancestry which originated in Ethiopia, and those populations are more genetically similar to the non-Africans, and may have experienced gene flow (Tishkoff, et al. 2009). So the non-African ancestry predicted in these populations could be through indirect contribution from their ancestral populations,” Wang et al. 2013, p. 7-8), they should’ve been able to detect it.
But the very fact that Wang et al. (2013) failed to detect a statistically significant fraction of Neandertal ancestry in San Bushmen is puzzling in and of itself considering that another recent study (not cited in Wang et al. 2013) reported West Eurasian admixture can be found all over South African foragers and pastoralists due to a pastoral migration from East Africa to South Africa in the past 3,000 years (see on the left, from Pickrell, Joseph K., Nick Patterson, Po-Ru Loh, Mark Lipson, Bonnie Berger, Mark Stoneking, Brigitte Pakendorf, and David Reich. 2013. “Ancient West Eurasian Ancestry in Southern and Eastern Africa”). Considering that ancient admixture follows the matreshka principle (“It should be noted that although ADMIXTURE predicts the contribution of three putative ancestral populations in each individual genome, the contribution could be indirect…MKK and Sandawe might not have large amounts of Eurasian ancestry. They have large amounts of Cushitic ancestry which originated in Ethiopia, and those populations are more genetically similar to the non-Africans, and may have experienced gene flow (Tishkoff, et al. 2009). So the non-African ancestry predicted in these populations could be through indirect contribution from their ancestral populations,” Wang et al. 2013, p. 7-8), they should’ve been able to detect it.
.



